Laboratory of Cancer Epigenetics
In the last years, the field of epigenetics has expanded into an essential component of biological and medical research, with modifications of DNA and histones being considered a center element in clinical oncology nowadays. For instance, drugs blocking DNA methylation have been approved by the FDA for the treatment of specific leukemic cancers.
Overall the laboratory of Cancer Epigenetics led by Prof. Fuks aims to investigate the regulation of epigenetic modifications during the processes of transcription and the mechanisms in which these processes are deregulated in cancers.
Besides studying fundamental mechanisms of cancer epigenetics (Part I), the group of F. Fuks developed a deep interest in cancer epigenomics to develop diagnostic and prognostic tools (Part II).
Completely unsuspected features were recently discovered that gave impetus to a new field, termed «Epitranscriptomics». In particular, the exciting potential of RNA modifications to fine-tune complex functions of mRNAs is revolutionizing our perception of epigenetics. In the last years, the Fuks’ lab shifted most of his interest to the nascent epitranscriptomic field (Part III).
Part I. Mechanisms of DNA modifications and their dysregulations in cancers.
In the past years, Prof. Fuks and his team addressed the mechanisms by which genes are regulated by DNA modifications (DNA methylation and hydroxymethylation/hmC). They revealed a much more direct connection than anticipated between DNA methylation and several histone modifications. Furthermore, they provided the first hints of how DNA methylation can be aberrantly targeted to specific loci in the genome during cancerogenesis. Several ongoing clinical trials stem from these studies.
Regarding DNA hydroxymethylation, the laboratory of F. Fuks highlighted a novel mechanism by which this modification promote transcriptional activation. More recently, using mouse cancer genetics, they highlighted the involvement of TET/hmC in tumour initiation and progression, and emphasizes the importance of epigenome plasticity in cancer development.
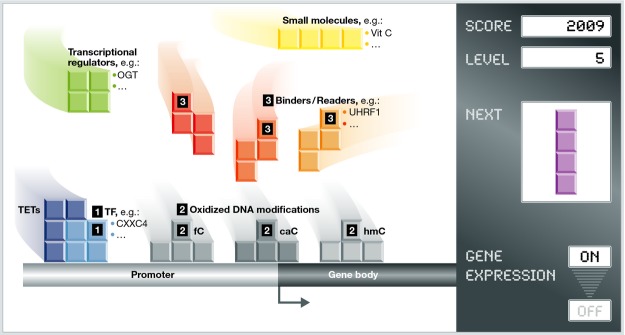
A picture is emerging in which TETs build up specific “DNA modification blocks” (hmC, fC, caC) through their coordinated regulation at multiple levels. The following sequence of events can therefore be suggested: (i) Transcription factors will result in (ii) 5hmC, 5fC, and 5caC formation with 5hmC being more abundant in gene bodies. (iii) These modifications can be further bound by various “binders” or “readers” that will either participate in DNA demethylation or translate the epigenetic signal to transcriptional activation or repression. Other active players are transcriptional regulators and small molecules such as vitamin C that directly affect TET activity and hence the deposition of 5hmC, 5fC, and 5caC. These DNA modification blocks, TETs, and TET regulators can assemble in a gene- and/or a cell-state-dependent manner creating precise landscapes which will ultimately affect gene expression.
Main Publications related to Part I :
- Dube et al. J Exp Clin Cancer Res 2023
- Bizet et al. Epigenetics 2022
- Bonvin et al. Cancer Research 2019
- Delatte et al. EMBO J. 2014
- Deplus et al. EMBO J. 2013
- Fuks F., Nature 2010
- Brenner et al. Dev. Cell 2007
- Villa et al. Cancer Cell 2007
- Brenner et al. EMBO J. 2005
- Hugues-Davies et al., Cell 2003
- Di Croce et al. Science 2002
- Fuks et al. Nature Genetics 2000
Part II. The Epigenomic of Cancer: the translational side
Large-scale cancer studies of the epigenetic landscape are reshaping our understanding of tumor heterogeneity and it is clear that epigenetic regulators are critical mediators of cancer progression. By acquiring and developing as soon as 2010 a NGS platform dedicated to epigenomics, the group of Prof. Fuks contributed significantly to the field of cancer epigenomics. His group indeed provided both novel biological insights and new avenues for cancer research. For example, the group has been one of the first to use the « Illumina Infinium Methylation » technology and performed the largest and most comprehensive DNA methylation profiling study of breast cancer, laying the ground for better understanding its heterogeneity and improving tumour taxonomy.
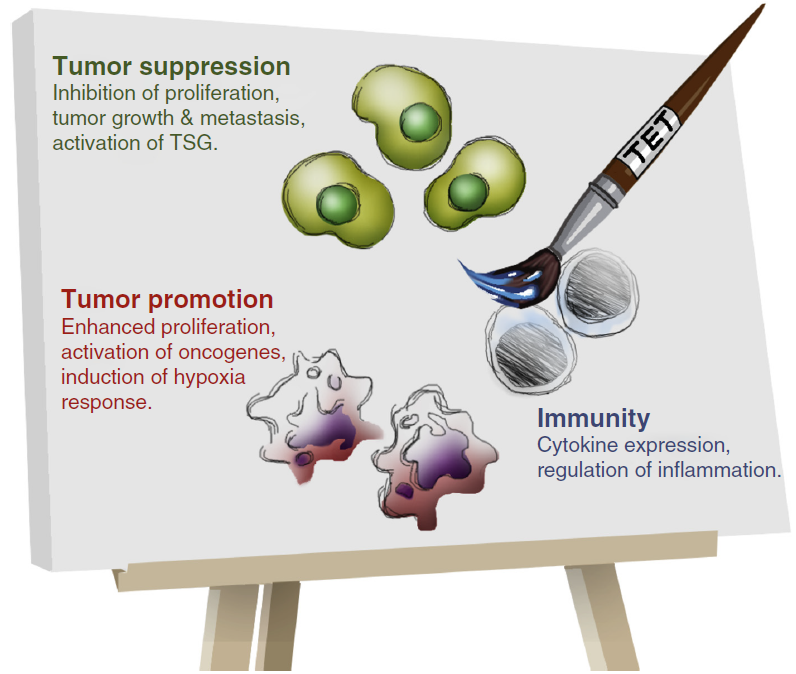
The lab also focused on developing diagnostic and prognostic tools. As cancer immunotherapy is rapidly advancing and gains significant public and scientific attention, the Fuks’ lab tackled the fundamental basis of epigenetics in the context of cancer immune response. In particular, the Fuks’ team has identified a specific DNA methylation signature that can predict which patients could benefit from immunotherapies.
Main Publications related to Part II :
- Collignon et al., Science Advances 2018
- Jeschke et al., Journal of Clinical Investigation 2017
- Van Grembergen et al., Science Advances 2016
- Volkmar et al. EMBO J. 2012
- Dedeurwaerder et al. EMBO Molecular Medicine 2011
- Ndlovu et al. Trends Biochem Sci. 2011
Part III. An emerging realm of biomedical research: Epitranscriptomics
Beside DNA methylation and histone modifications, there is an increasing interest in RNA modifications. All in all, knowledge about the physiopathological impacts of epitranscriptomic marks should change the way we think about gene regulation in health and disease.
- Only few mRNA modifications are known to date. In a seminal work leading to breakthrough discoveries, the Fuks’ lab recently addressed the distribution, function and biological role of a novel RNA modification (RNA hydroxymethylation). In a follow up study, the team deciphered the physiological importance of this new modification in embryonic stem cells.
- One of the most outstanding question in the booming field of RNA epigenetics is to evaluate the contribution of epitranscriptomics to cancer. Recently, the team of F. Fuks set up a comprehensive framework to study the dysregulation in cancer of the most prevalent mRNA modification, m6A. They reveal an unexpected mechanistic link between an mRNA mark and key cancer processes. They also highlighted m6A as a prognostic marker and a stratification guide for patient-tailored cancer therapy.
- Using cutting-edge technologies (e.g. chemical biology, NGS and RNA mass spectrometry) as well as up-to-date model systems (e.g. mouse PDX), ongoing studies in Prof. Fuks’ lab are addressing the most burning questions of the field, such as :
- 1 Characterization of novel RNA modifications and enzymes
- 2 Physiopathological impacts of epitranscriptomics
- 3 Clinical applications of RNA modifications
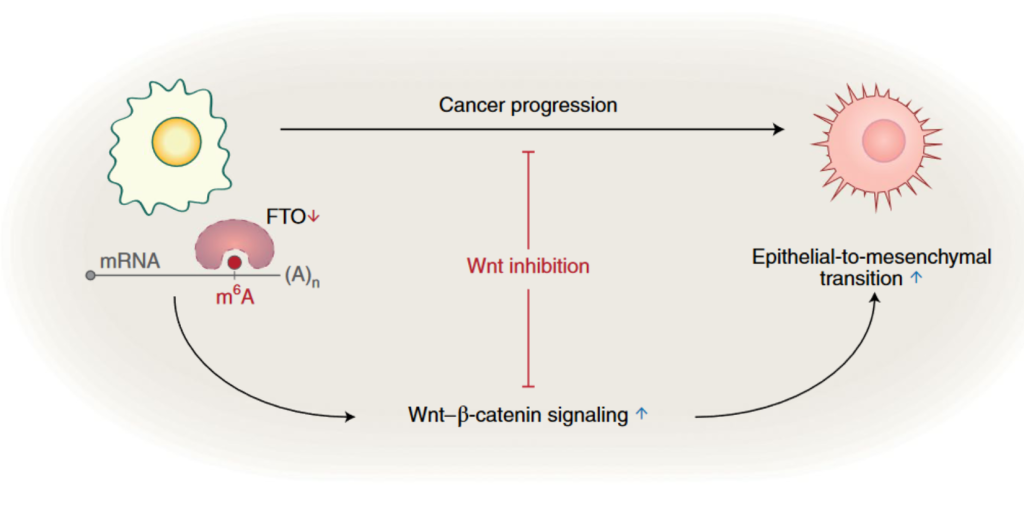
Downregulation of FTO (red downward arrow) in epithelial tumors leads to an increase in m6A modification on mRNAs, which then results in activation of the Wnt-ß-catenin signaling pathway (blue upward arrow). Wnt-ß-catenin acts upstream of EMT to promote cancer progression. The study by Jeschke et al. proposes the use of Wnt inhibitors as a therapeutic strategy to target FTO-low tumors.
Main Publications related to Part III :